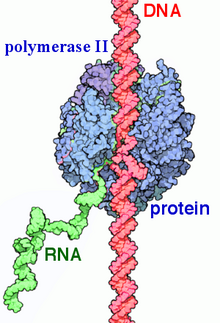
Phosphorylation-Regulated Binding of RNA Polymerase II to Fibrous Polymers of Low-Complexity Domains: Cell

Roles of DNA polymerases V and II in SOS-induced error-prone and error-free repair in Escherichia coli | PNAS

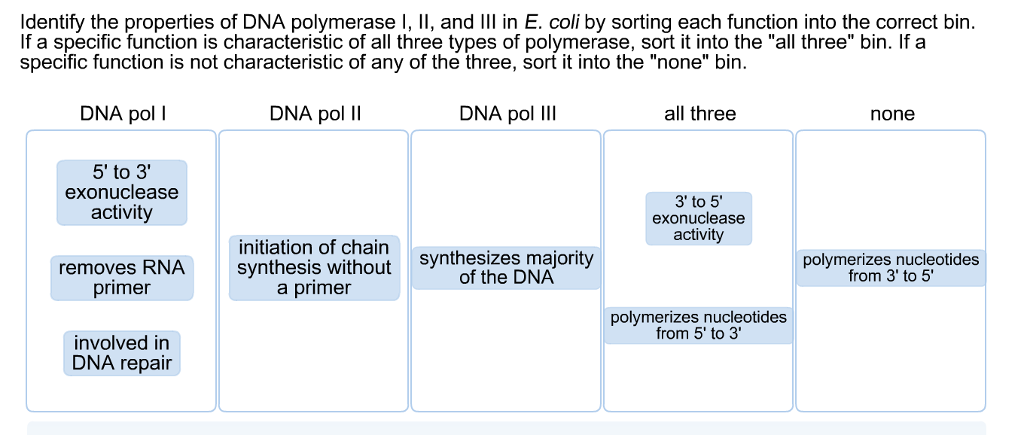
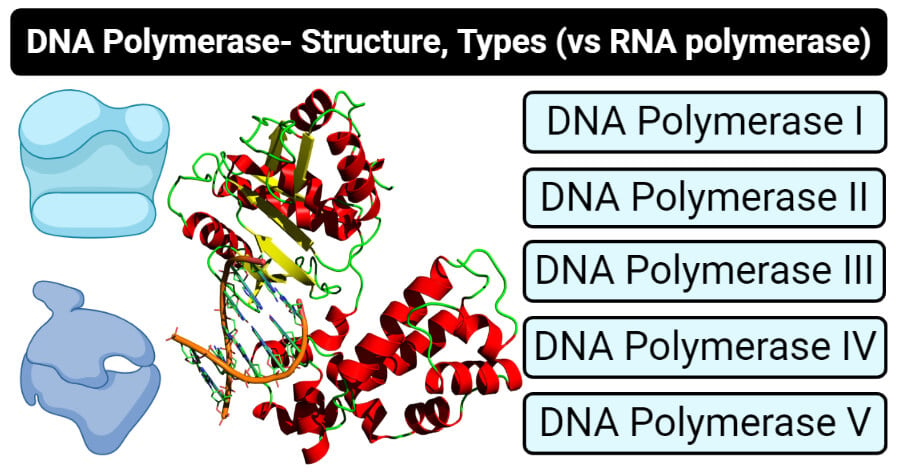
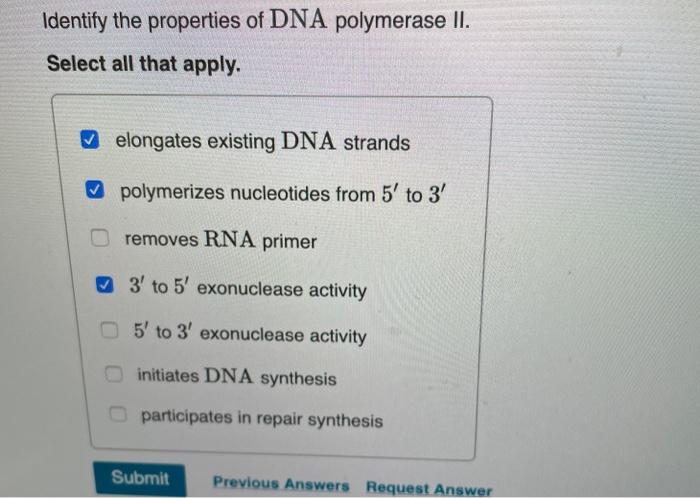
SOLVED: 17 State the function of three types of DNA polymerases: 1. DNA polymerase 1 (DNA Pol I), 2 DNA polymerase 2 (DNA Pol II); and 3 DNA polyinerase 3 (DNA Pol III): 8 (3 Points)
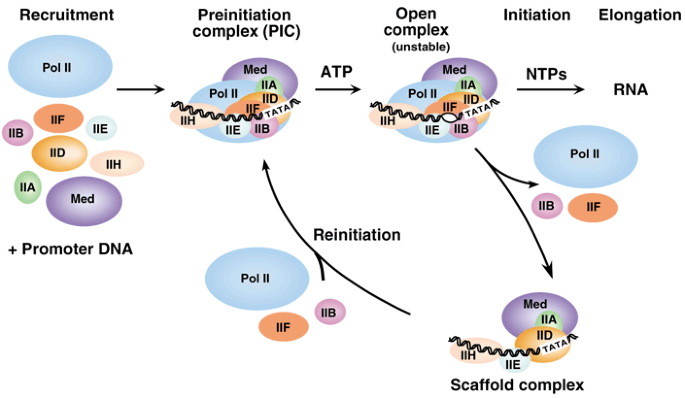
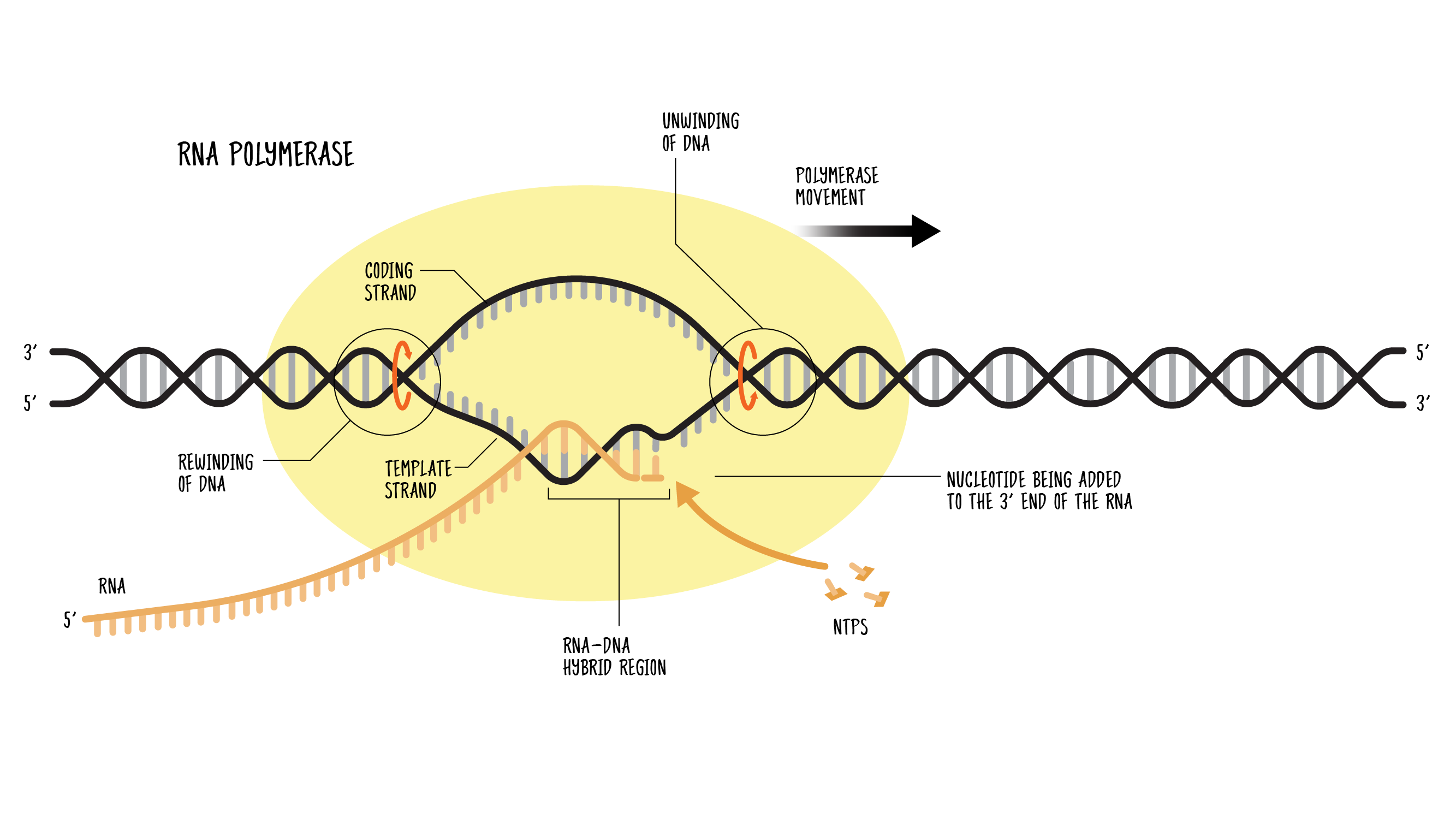
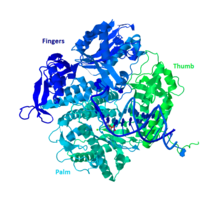
Structure and mechanism of the RNA polymerase II transcription machinery | Nature Structural & Molecular Biology

Stimulation of RNA Polymerase II ubiquitination and degradation by yeast mRNA 3′-end processing factors is a conserved DNA damage response in eukaryotes - ScienceDirect

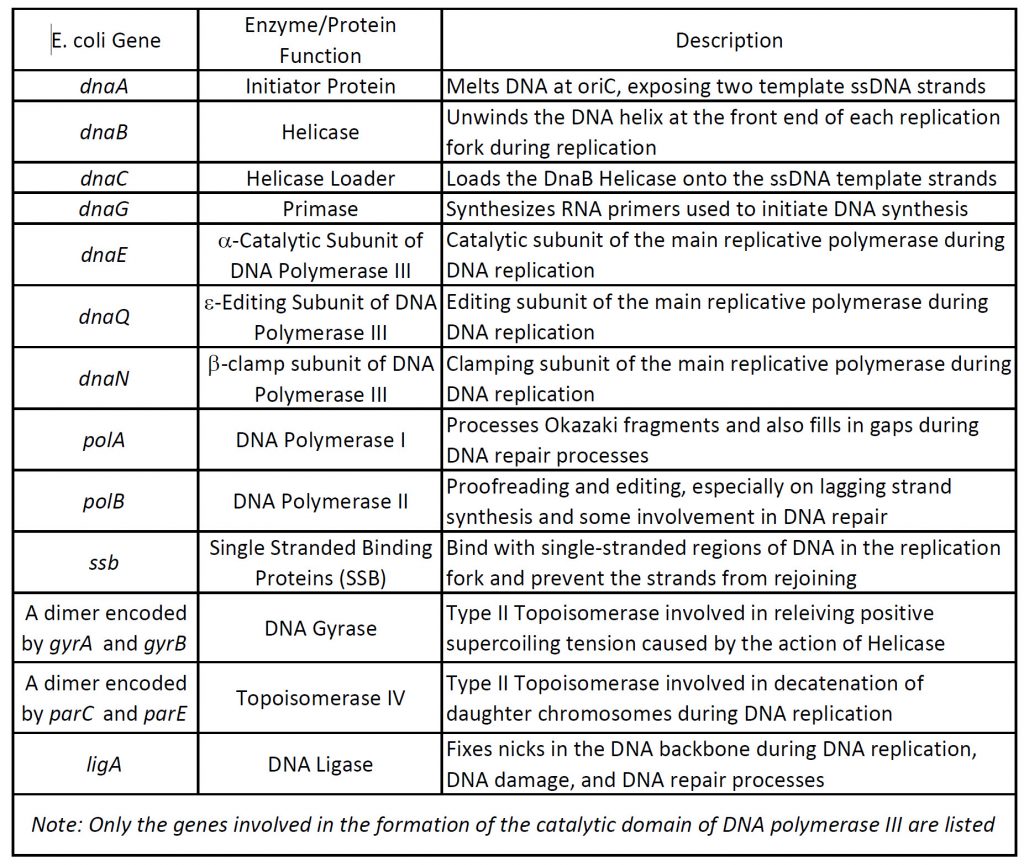
Regulation of fidelity of DNA replication in Escherichia coli and Saccharomyces cerevisiae cells | MolBioPhd
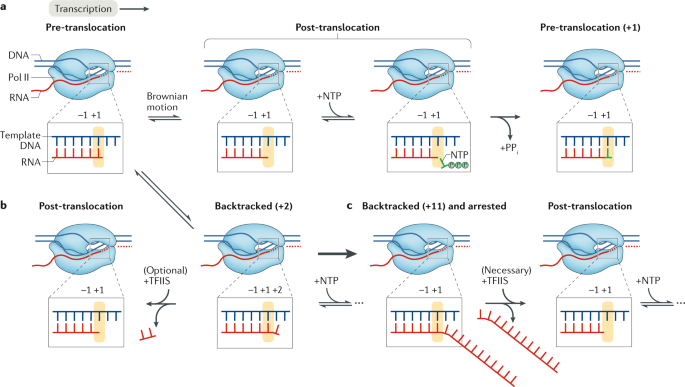
Causes and consequences of RNA polymerase II stalling during transcript elongation | Nature Reviews Molecular Cell Biology